Universal Data Structure (UDS)
Universal Data Structure (UDS) aims to provide a unifying framework on how to represent tree/graph network data displayed & modified in the frontend and backend. It aims to unify the data representation used to communicate between both ends, providing seamless transition between services.
Data Structure Trinity
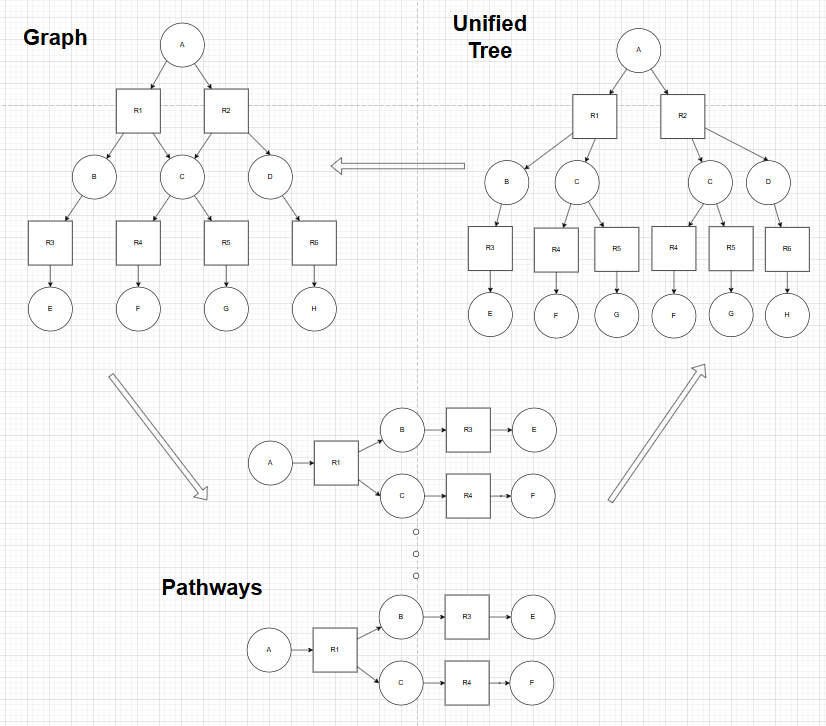
There are 3 major data structures which collectively represent the metadata & connectivity of synthesis pathways in ASKCOS. They are known as Graph, Unified Tree & Pathways. Typically, a Tree Builder (TB) job would be working either on a Graph (MCTS), or Unified Tree (Retro*) data structure when searching for synthesis pathways.
Further post-processing on the Tree Builder job yields a complete acyclic graph which captures the entirety of the searched space, and enumeration of the search graph produces individual pathways. The Tree Builder job can be viewed in 2 ways, one through Pathways data structure in Tree Explorer (TE), another through Unified Tree data structure in IPP. On the other hand, an expand one call from the frontend IPP page interacts with the backend, modifying the IPP on the fly.
It is noteworthy to point out, modification done on the canvas of IPP and Tree Explorer inherently modifies the frontend dispGraph and dataGraph.
Graph
- Directed Acyclic Graph
- Node ID: SMILES
- Used in TB graph output, IPP/TE
dataGraph
Unified Tree
- Directed Rooted Trees
- Node ID: UUID
- Used in IPP/TE
dispGraph
Pathways
- Multiple Directed Rooted Trees
- Node ID: UUID
- Used in TB path output
UDS representation format
To unify the 3 different data structure, UDS is designed to capture all nuances of this representation to eliminate redundancy in representation and facilitate easier editing through a more flattened representation.
{
"uds": {
"node_dict": {
"CHEM_SMILES_1": {
CHEM_SMILES_1_METADATA
},
"CHEM_SMILES_2": {
CHEM_SMILES_2_METADATA
},
"RXN_SMILES_1": {
RXN_SMILES_1_METADATA
},
"RXN_SMILES_2": {
RXN_SMILES_2_METADATA
},
...
},
"graph": [
{
"source": CHEM_SMILES_1,
"target": RXN_SMILES_1,
},
{
"source": CHEM_SMILES_2,
"target": RXN_SMILES_2,
},
...
],
"uuid2smiles": {
UUID_1: CHEM_SMILES_1,
UUID_2: CHEM_SMILES_2,
UUID_3: RXN_SMILES_1,
UUID_4: RXN_SMILES_2,
...
},
"pathways": [
[
{
"source": UUID_1,
"target": UUID_2,
},
{
"source": UUID_3,
"target": UUID_4,
},
...
],
[
{
"source": UUID_5,
"target": UUID_6,
},
{
"source": UUID_7,
"target": UUID_8,
},
...
],
...
],
"pathways_properties": [
{
PATH_PROP_1
},
{
PATH_PROP_2
},
...
]
}
}The representation format above consists of node_dict , graph , uuid2smiles , pathways ,pathways_properties section. It separates the node metadata section and the connectivity to remove redundancy in representation, allowing easier modification of information. Connectivity of graph/pathways is stored in a nodelink format.
node_dict - Stores all of the metadata of chemical node or reaction node returned in the expand-one call. Dict of Dict
graph - Stores the search graph connectivity in nodelink format. List of Dict
uuid2smiles - Mapping of UUID to SMILES, facilitating reconstruction of UDS back to Graph Object. Dict
pathways - Stores the pathways connectivity in nodelink format. List of List of Dict
pathways_properties - Store the pathways properties from tree analysis jobs. List of Dict
Chemical Node Dictionary
{
"smiles": "CN(C)CCOC(c1ccccc1)c1ccccc1",
"as_reactant": 59,
"as_product": 44,
"properties": [
properties_list
],
"purchase_price": 4.13,
"terminal": false,
"type": "chemical",
"id": "CN(C)CCOC(c1ccccc1)c1ccccc1"
}Reaction Node Dictionary
{
"smiles": "CN(C)CCCl.OC(c1ccccc1)c1ccccc1>>CN(C)CCOC(c1ccccc1)c1ccccc1",
"precursor_rank": 1, // rank from reranker
"precursor_score": -0.003586091330295326, // score from reranker
"plausibility": 0.9981883764266968,
"rxn_score_from_model": 0.334626168012619, // # average of normalized_model_score
"model_metadata": [
{
"direction": "retro",
"backend": "template_relevance",
"model_name": "reaxys",
"attributes": {
"max_num_templates": 1000,
"max_cum_prob": 0.995,
"attribute_filter": []
},
"model_score": 0.334626168012619,
"normalized_model_score": 0.334626168012619,
"rank": 1,
"reaction_id": null,
"reaction_set": null,
"source": {
"template": { ... }, // template relevance
"reaction_data": {} // retrosim
}
},
{
"direction": "retro",
"backend": "augmented_transformer",
"model_name": "USPTO_FULL",
"attributes": {},
"model_score": 0.13734960132034285,
"normalized_model_score": 0.17782974596553017,
"rank": 2,
"reaction_id": null,
"reaction_set": null,
"source": {
"template": null,
"reaction_data": null
}
}
],
"precursor_properties": {
"rms_molwt": 150.57960055284425,
"num_rings": 2,
"scscore": 1.51275690065044
},
"reaction_properties": {
"canonical_reaction_smiles": "CN(C)CCCl.OC(c1ccccc1)c1ccccc1>>CN(C)CCOC(c1ccccc1)c1ccccc1",
"mapped_smiles": "Cl[CH2:5][CH2:4][N:2]([CH3:1])[CH3:3].[OH:6][CH:7]([c:8]1[cH:9][cH:10][cH:11][cH:12][cH:13]1)[c:14]1[cH:15][cH:16][cH:17][cH:18][cH:19]1>>[CH3:1][N:2]([CH3:3])[CH2:4][CH2:5][O:6][CH:7]([c:8]1[cH:9][cH:10][cH:11][cH:12][cH:13]1)[c:14]1[cH:15][cH:16][cH:17][cH:18][cH:19]1",
"plausibility": 0.9981883764266968,
"reacting_atoms": [
5,
6
],
"selec_error": null,
"cluster_id": null,
"cluster_name": null
},
"type": "reaction",
"id": "CN(C)CCCl.OC(c1ccccc1)c1ccccc1>>CN(C)CCOC(c1ccccc1)c1ccccc1"
}